Decoding the Genetic Code - Awe and Reverence
Wonders, Amazement and Delightful Puzzles (UNDER CONSTRUCTION 8/29/2022.)
The three previous posts have attempted to at least describe (even if not completely explain) the wonders of the Genetic Code. Along the way of discovery, I encountered some delightful puzzles that completely defy explanation. The old “The xxx-protein evolved the ability to perform …miracles…” just isn’t Science. In this post I am going to dive into some of these questions.
I am a retired electronics engineer. I love to “reverse engineer” challenging products or designs that I encounter. I just want to understand how things work. Sometimes the best way to understand some functions is to figure out (at least in principle) how I would “design” this function, at least in my imagination. This allows me to gain insights into the basic nature of the problem. “What needs to be accomplished to achieve the observed end?”
All of these “solutions” require a broad overview of the system. Otherwise, a “solution” within one system will only create new problems in other systems. AN “ideal” solution should aim to solve problems across multiple systems. The “real-life” solutions clearly imply a huge and profound amount of advance planning and insight — both of which are forbidden in Darwinian Theory. Most of these are “Chicken and Egg” paradoxes. Occasionally, someone will come up with an unexpected resolution to one or two “Chicken and Egg” paradoxes. But there are literally hundreds. Over time I have learned to treat these with with reverence, and just enjoy them.
During the Human Genome Project (1990-2003) the aim was to sequence the whole human genome. Planning started after the idea was picked up in 1984 by the US government, the project formally launched in 1990, and was declared essentially complete on April 14, 2003, but included only about 85% of the genome. Level "complete genome" was achieved in May 2021, with a remaining only 0.3% bases covered by potential issues. The final gapless assembly was finished in January 2022 (en.wikipedia.org/wiki/Human_Genome_Project). The early emphasis was on locating and identifying our genes. Genes are the specific segments of DNA that contain the instructions for assembling a protein. As I recall, they initially estimated that our genes occupied about 25% of the genome. In time, that dropped to 15%. Then 10%. Then 5%. Finally, a reporter asked the obvious question, “Then what is the other 95%?” The answer stunned me. “Oh, that is just ‘junk DNA’: the leftover scratch paper of Evolution.” After all, the “genes” are just the instructions for assembling proteins. “Making” all 20,000 genes would only get you some rich protein soup — not a living organism. The genes are only the “Parts List” for the organism. Where then is the “Assembly Manual”? Where is the “Operating Manual”, or the “Repair Manual”? It was obvious even back then that the science was still in its infancy.
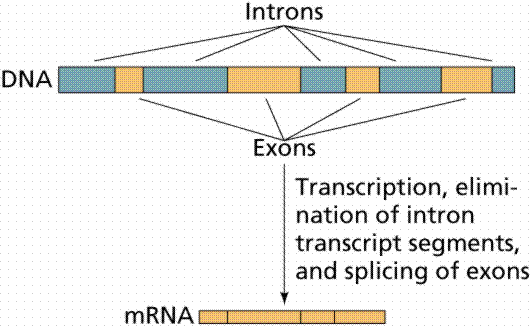
It was once thought that the coding in a gene had a 1:1 relationship to the finished product. It was soon discovered that the gene seemed rather fragmented. There were “non-functional” sections called "introns" interspersed throughout the gene. After the gene is transcribed by RNA Polymerase into a strand of messenger RNA (mRNA), these “superfluous” sections must be snipped out, and the gaps spliced together[1]. This is performed by spliceosomes[2].
This in itself is a startling discovery. This seems to add a lot of "unnecessary" complexity (with many potential errors) to the transcription process. But explaining the origin of this gene splicing is particularly problematic. First, there has to be an overall plan. Second, there must be an organized procedure for implementation. Third, it requires that there must be a group of specialized protein machines which can carry out some complex and critical operations[3]. Fourth, these all must come together and work flawlessly. The entire evolutionary process could be completely derailed at any stage of the evolution of gene splicing that fails to mature. This scheme has to work the first time.
[1] p 103
[2] These are spliceosomal introns because even prokaryotes actually do have a certain type of introns. They are related to eukaryotic introns called "group I and II' introns found in tRNA and rRNA . The RNA produced from these introns have their own special activity- they can splice themselves. Sep 10, 2009 http://genetics.thetech.org/ask/ask326
[3] https://www.dnalc.org/view/16933-3D-Animation-of-DNA-to-RNA-to-Protein.html (an animated movie)
It was initially unclear whether these introns were simply useless "filler material", but they actually contain additional instructions and functions soon to be discovered which guide and direct the splicing operations.
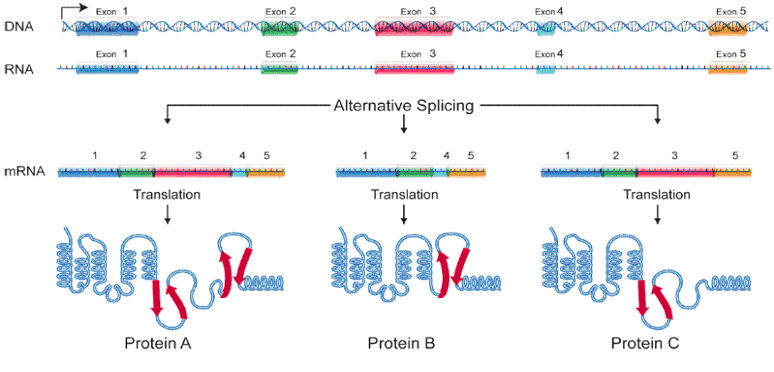
But it gets even more complex. Researchers discovered that there are actually overlapping genes. This is accomplished as shown below. Here, three hypothetical proteins share the same "framework" structure that supports the active site of each protein. By selecting which "optional" exons will be copied, several different "final products" can be stored in the same gene. The less-critical supporting structure design is re-used for making several different proteins. In other cases, this supporting structure has to be "bent" slightly to put pressure on the active site to fine-tune it for absolute maximum performance. Such a protein cannot use a generic "stock" framework unless alternate "modification exons" are specified. Think for example of a “stock” screwdriver handle and a selection of 6 “optional” screwdriver blades.
Only after selective splicing is the mRNA ready for production. The alpha helix and other structures will fold into their final three-dimensional shape to complete the process.
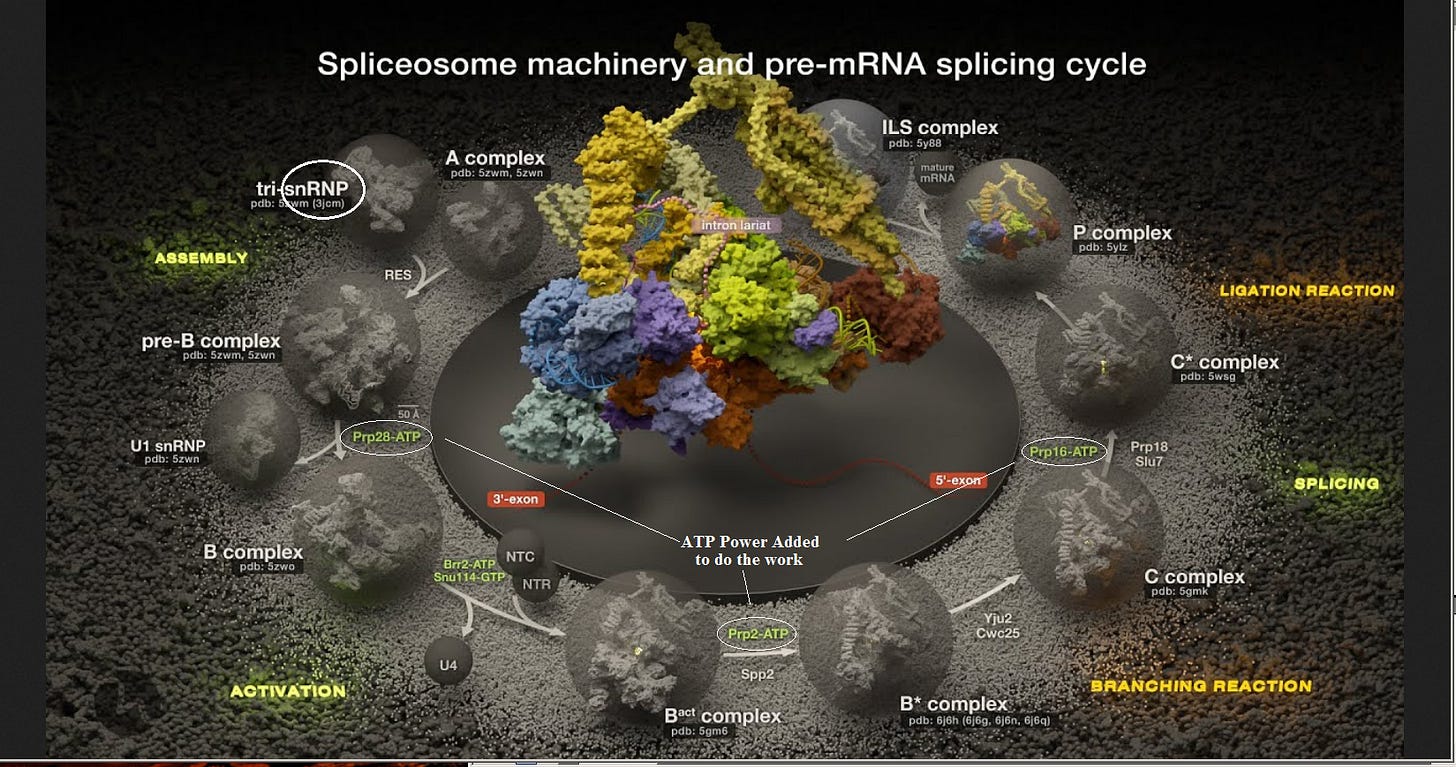
This is fascinating… A spliceosome arrives with a “work order” for Protein C. Are the instructions for Exon Splicing in the Introns? Does it “remember” the instructions before discarding the Introns? Interesting question! But further intron questions come up later…
Just consider the “snRNP” (upper left) at the start of the cycle. “snRNPs (pronounced "snurps"), or small nuclear ribonucleoproteins, are RNA-protein complexes that combine with unmodified pre-mRNA and various other proteins to form a spliceosome, a large RNA-protein molecular complex upon which splicing of pre-mRNA occurs. The action of snRNPs is essential to the removal of introns from pre-mRNA, a critical aspect of post-transcriptional modification of RNA, occurring only in the nucleus of eukaryotic cells. Additionally, U7 snRNP is not involved in splicing at all, as U7 snRNP is responsible for processing the 3′ stem-loop of histone pre-mRNA. [1]
The two essential components of snRNPs are protein molecules and RNA. The RNA found within each snRNP particle is known as small nuclear RNA, or snRNA, and is usually about 150 nucleotides in length. The snRNA component of the snRNP gives specificity to individual introns by "recognizing" the sequences of critical splicing signals at the 5' and 3' ends and branch site of introns. The snRNA in snRNPs is similar to ribosomal RNA in that it directly incorporates both an enzymatic and a structural role.” (en.wikipedia.org/wiki/SnRNP).
So the specific instructions as to how the spliceosome should delete introns and select which exons are to be included are included in the snRNA.
This is yet another clear example of “Irreducible Complexity”. During the splicing procedure, a the spliceosome undegoes a whole series of “shape ransfomations” that perfom the highly complex task of grabbing the ends of the adjacent exons to be preserved, snipping out and discarding the intron, then pulling the loose ends together and splicing them.
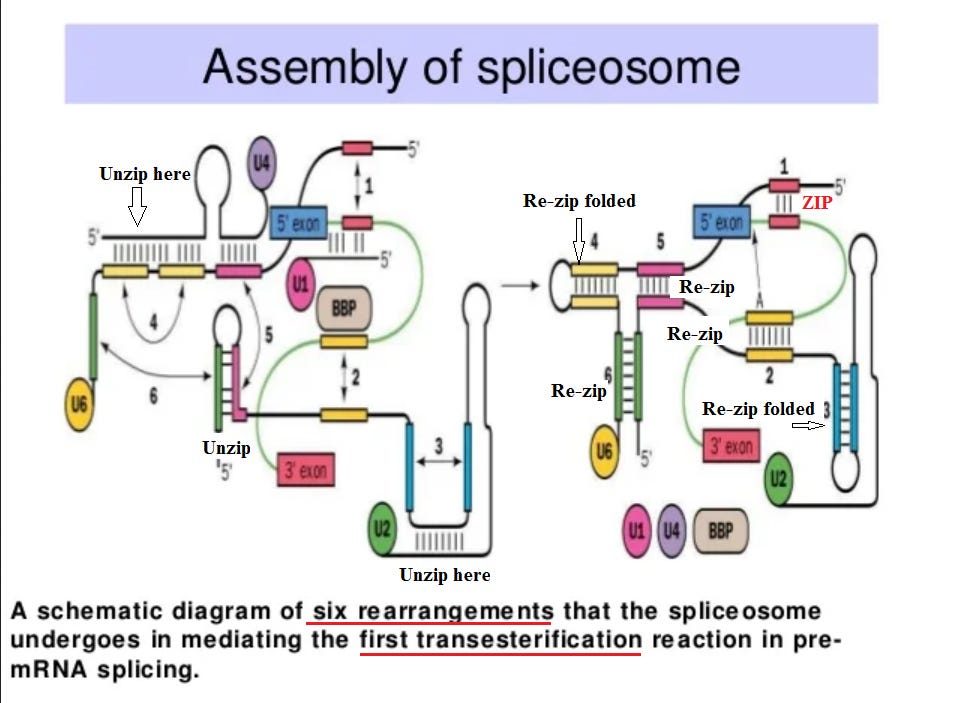
But how does a spliceosome perform its task successfully? This is fascinating! Simply to perform the “first transesterification”, it goes through six shape changes (like a toy “transformer”).
There are in this diagram what appear to be “zippers”, and that analogy may be useful. Weak bonds between adjacent strings of amino acids hold them together in the overall folded structure. The strength of the zipper can be precisely adjusted by changing the length. When the spliceosome binds to the pre-RNA at an intron juncture, that places a strain on the structure. This is just enough to trigger an unzipping at position (1), and the amino acid strings “re-zip” in a different position. This triggers a whole series of shape changes. Note in the previous figure that there are three points in the cycle where energy must be added in the form of ATP (adenosine tri-phosphate), which is the cell’s standard form of energy “fuel”. Again, “simply” to perform the “first transesterification”, it goes through six shape changes (like a toy “transformer”). In the previous figure, there are a whole series of additional steps involved.
This whole process seems problematic. One would certainly assume that the earliest cells would use a simple “1:1” RNA mapping, and the spliceosome would not even be needed. Actually adding the spliceosome function adds what seems to be an unnecessarily complex and dangerous new procedure. It is ripe for errors. This procedure is also “Irreducibly Complex”. ALL of the spliceosome complex parts must suddenly appear simultaneously and work correctly. If even one is missing or is not properly functional, then there is a very high probability that the whole splicing function will fail.
As I said, I am a retired engineer. When I am “reverse engineering” a product or a design and I come across some extremely complex features, I have to assume that there must be a very serious problem that required an extreme “fix”. But I don’t see any serious problem here that would justify such complexity. Further down the road I found some fascinating problems which might possibly account for introns. These provide some wonderful rabbit holes to chase down!
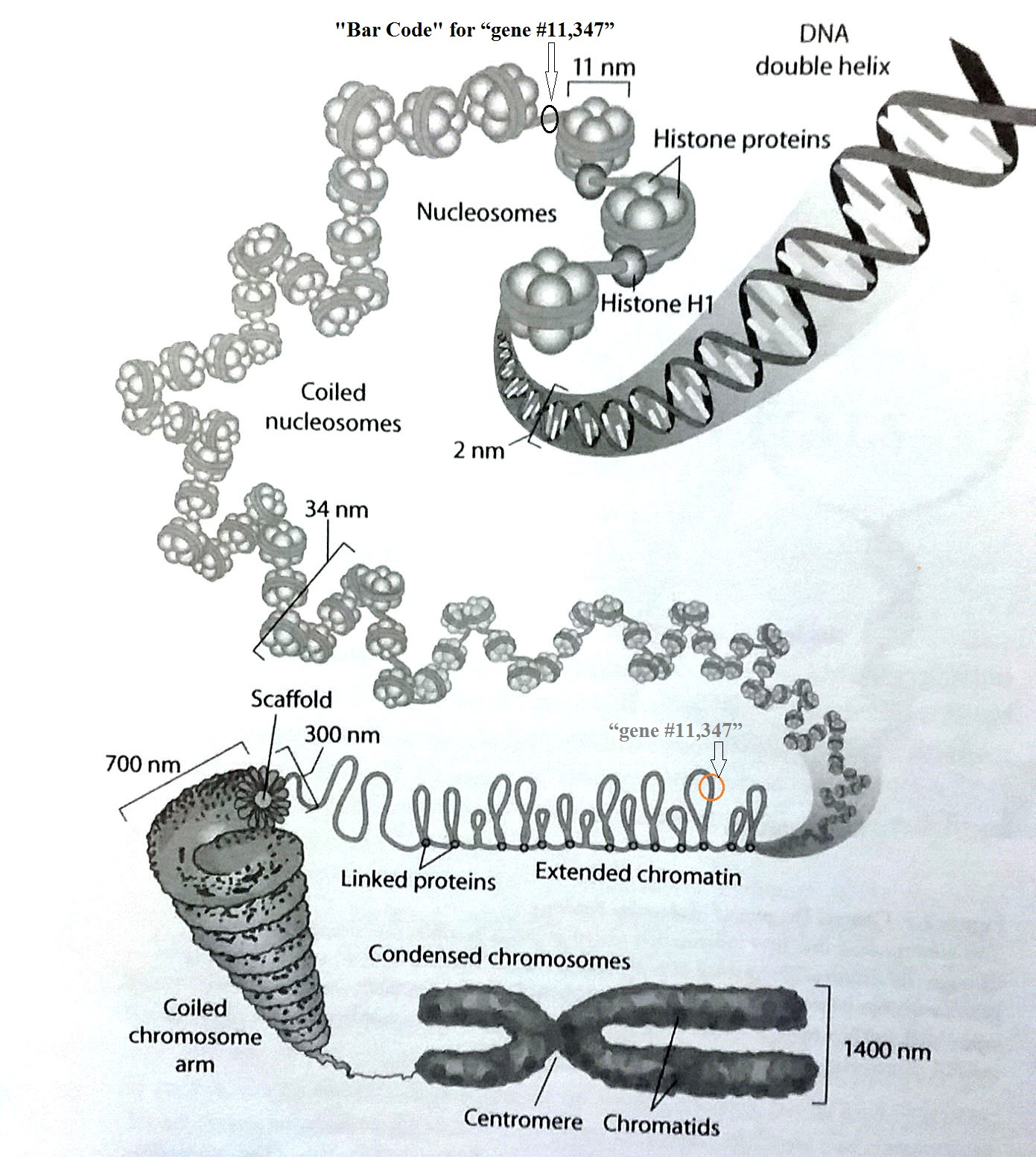
This figure is from Part 2: (the search for “gene #11,347”)
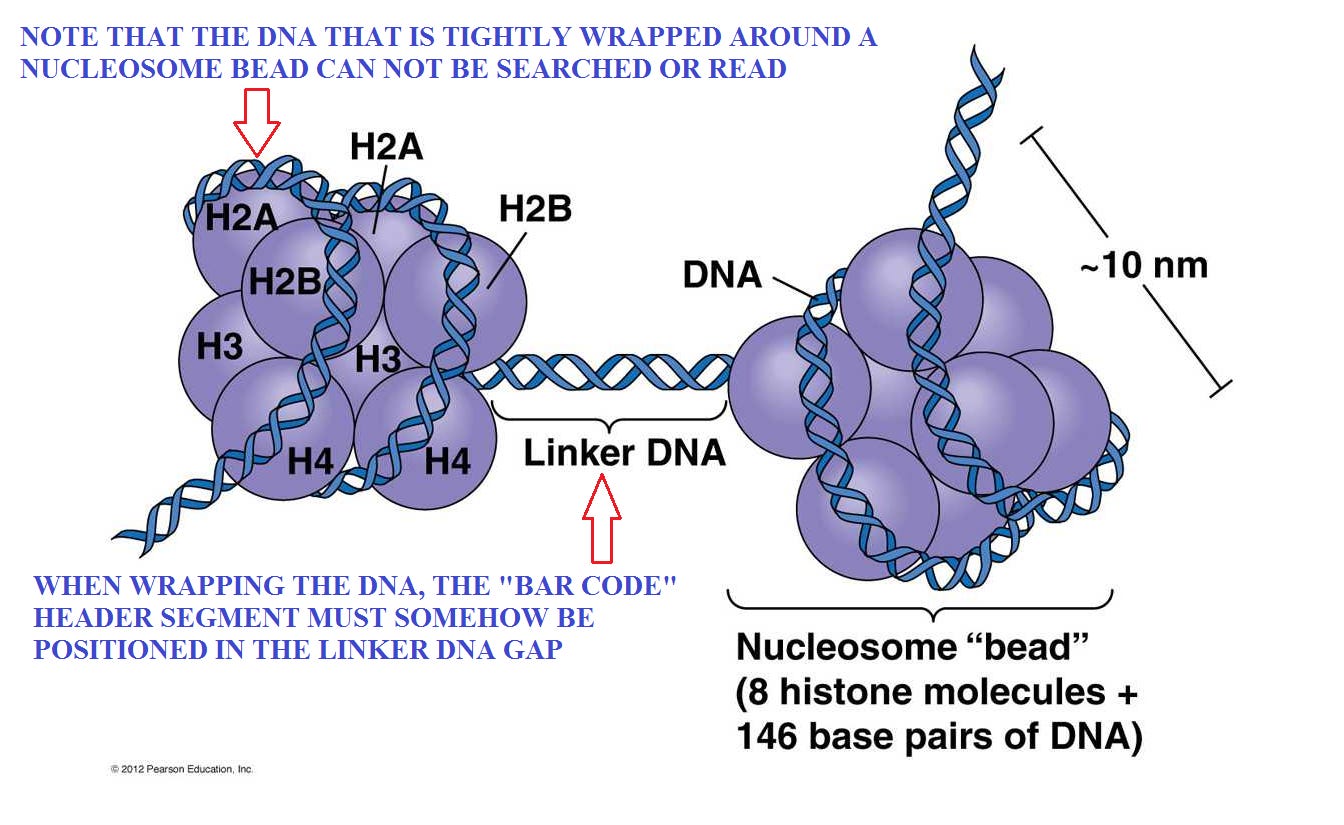
The header “bar code” for that needed gene must be accessible and visible.
The size, shape and spooling capacity of the nucleosome bead is fixed. So that implies that it may be necessary to make some adjustents to the length of the DNA to ensure that the proper and important code appears in the accessible linker segments. Otherwise, some large adjustments in the lengths of linker segments might be required. Also, recall that the genes are only a few percent of the total length, so they provide very limited flexibiliy. So while introns may play some part, we need to find some more robust possibilities.
Going back to Part 1, “On the average any amino acid can be "spelled" three different ways. This is done by simply substituting one of the alternate codes that specify the same amino acid. This provides a tremendous flexibility in "spelling" a protein. A protein with a length of 100 residues can be "spelled" roughly 3^100 ways. That is 5 x 10^47 ways to "spell" the same protein. (500,000,000,000,000,000, 000,000,000,000,000,000,000,000,000,000 ways!) But even any segment as short as 12 base pairs (4 amino acids) can be "spelled" about 81 ways.”
Imagine if we had that sort of flexibility in spelling. That would make it possible to impose a spelling pattern that required every 12th letter to be an ‘a’. Now it might be possible to force the DNA to just drop onto the nucleosomes “like a bicycle chain on a sprocket wheel.” This is just speculation, and vastly oversimplified, but it suggests that the problem is surely not impossible. Something like this seems to be happening.
Your genome contains genes for over 700 amazing proteins that spend their full time just reading your DNA and looking for errors.
Fixing Source Code Errors
During the replication of your DNA prior to cell division, errors sometimes occur. Considering the huge number of base-pairs (some 3.1 billion), a substantial number of errors are inevitable. To understand better why errors are "inevitable", just watch the process in real time:
http://www.hhmi.org/biointeractive/dna-replication-basic-detail
There is a whole suite of proteins that come to the aid of DNA Synthetase to perform a bewilderingly complex task at what seems like a break-neck speed. Note that this video is in "real time". They sometimes refer to DNA as being "self-replicating". This is absolutely NOT true. It takes a small army of highly complex proteins to perform extremely complex operations at dizzying speed to manufacture the duplicated DNA. But of course, errors DO occur.
This leads to another marvel that is still not well understood. There is error correction at work on the "source code": your DNA itself. A mutation in the DNA not only affects the next protein to be manufactured, but all subsequent copies as well. One research group has discovered over 700 proteins whose function is to traverse the length of your genome looking for errors. When it finds an error, it snips out that segment and corrects the error. 700 proteins is a major, significant fraction (3.5%) of your 20,000 proteins. Now obviously these “mere 700 proteins” can't "remember" my whole genome to compare and find specific errors. That would at least double the size of the genome if every cell carried "proof copies"! Even so, there could just as likely be errors in the “proof copy”! The Apollo Space Missions carried THREE computers, so that in case of a computer glitch, they could take a “majority vote”! Then you would need the “machinery” to compare the three separate DNA copies and correct the odd one. That seems like a losing cause!
It is suspected that the flexibility in "spelling" of protein sequences allows for the definition of "acceptable" and "unacceptable" longer sequences. If the gene coding selectively avoids “forbidden sequences” then the "Mr. Fix-It" enzymes (yes, that is what the web site actually originally named them ;>) do not need to know the specific "original code", but needs only correct "unacceptable pattern" errors. This is much like parity coding or the error metric used in FEC (Forward Error Correction coding) in high speed digital transmissions. Thus there is evidently a high level of quality control at work at every level of the cell. But that implies a “higher level supervisory intelligence”. This is another conundrum for evolutionist!
It is one thing to discuss and describe these high-level functions. But implementing them in tiny "dumb" proteins is a whole other story. These are tiny protein complexes which bind to the DNA double helix. They then "read" (by Braille, perhaps? ;>) the DNA code. They are able to "recognize" improper patterns and react to them by performing delicate and complex surgery. In principle, we could build highly complex computer-controlled robots which could be programmed to perform equivalent functions, (on a larger scale, or course!) but even that would require an immense amount of intelligence and intelligent planning.
THIS PAGE IS STILL A WORK IN PROGRESS…