Decoding the Genetic Code - Part 2
Gene Transcription -- Far more than just "copying"! Edited August 18, 2022
In Part 1 I explored the structure of DNA itself. Instructions for building every protein in my body are encoded in the “genes”. Three-letter “codons” specify the sequence of amino acids (there are only 20 amino acids actually used). There is a built-in redundancy (60 codes to specify just 20 amino acids), and the codon table is highly optimized to minimize errors and also to minimize the level of damage caused by errors. This implies either a very large degree of evolution, or a high degree of intelligent planning. But in nearly 70 years, no one has been able to suggest a remotely plausible way for even rather minor evolution of The Code to occur.
In this section, I will discuss the process of copying genes into messenger RNA (mRNA), pre-processing that mRNA, and transcribing it into actual proteins. The illustrations that I use in this section should not be taken literally. The concepts discussed here are extremely technical and complex, so I attempted to find familiar functions which help to visualize the end results. It has been my experience that when I get deeper I get into the study of a particular function, it invariably turns out to be vastly more mind-boggling than I anticipated. My wish is that these will help achieve a”general idea” of how things work and enable you to grasp the more complex ideas more readily when you encounter them.
There are some real mysteries when you start examining the process of gene transcription. Transcription is the first step of gene expression, in which a particular segment of DNA (a gene) is copied into RNA by the enzyme RNA polymerase. The messenger RNA (mRNA) is transported out of the nucleus where it is taken up by a ribosome and multiple proteins are assembled.
But before you can copy it, you have to find it. Your genome is over 3,100,000,000 base pairs long. When you cell needs to fabricate some more of a specific protein, it first has to locate the specific gene to be copied.
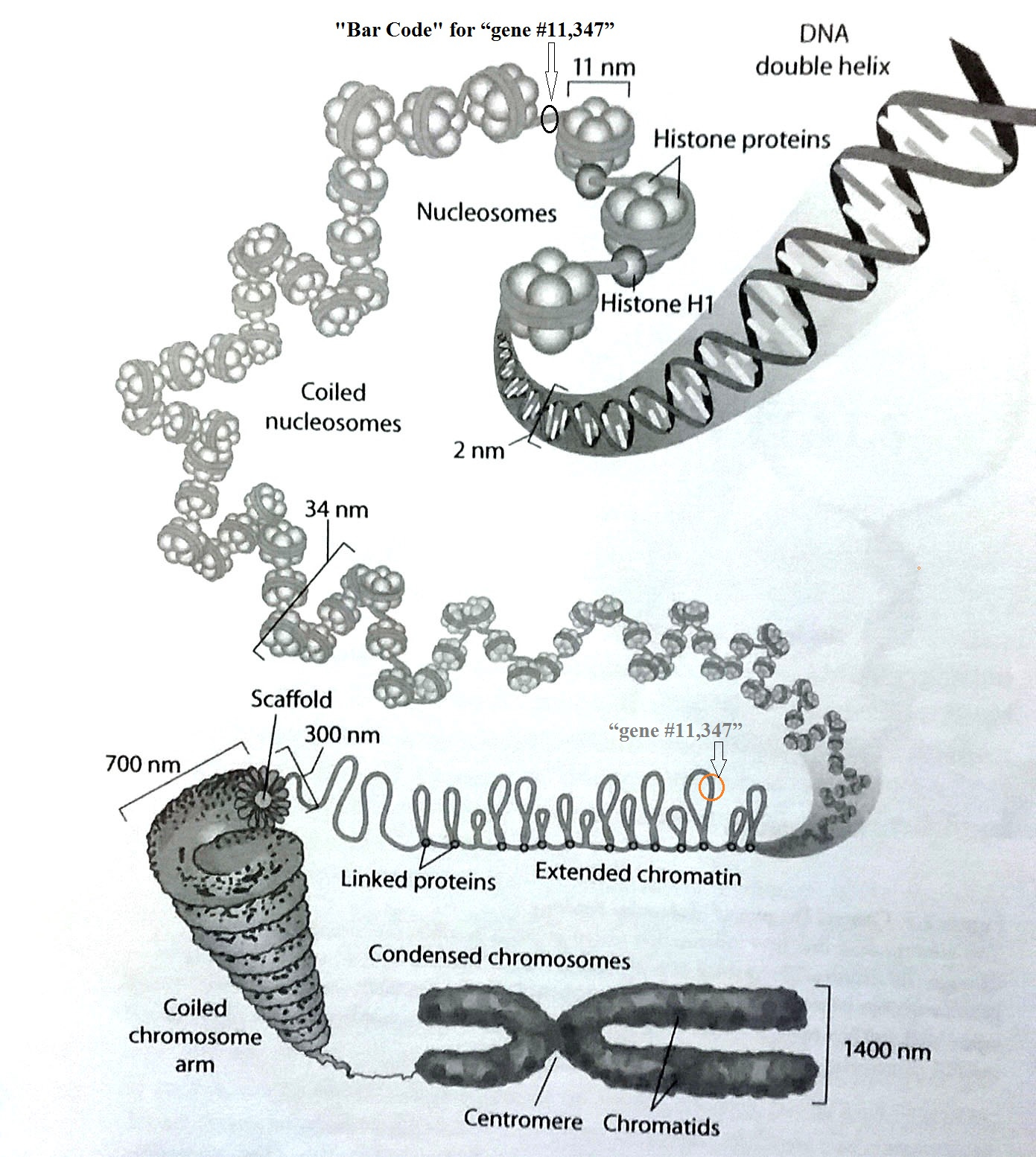
Figure 1. DNA-packing within a single chromosome. First the cell must locate the chromatin loop containing “#11,347” (middle) then find “Barcode 11,347”. The DNA must be uncoiled from the adjacent histone “spools” to make the whole gene accessible. (top)
Note in this figure that the “bare DNA” has a diameter of only 2 nanometers. That is 0.000,000,002 meters, or 0.000,000,080 inch. At the other end of the size spectrum is the typical chromosome, which is about 1400 nanometers (0.000,055 inch). That is 700x larger. There are 23 such chromosomes.
I add here an summary quote from https://www.sciencefocus.com/the-human-body/how-long-is-your-dna/
“The DNA in your cells is packaged into 46 chromosomes in the nucleus. As well as being a naturally helical molecule, DNA is supercoiled using enzymes so that it takes up less space.
Try holding a piece of string at one end, and twisting the other. As you add twist, the string creates coils of coils; and eventually, coils of coils of coils. Your DNA is arranged as a coil of coils of coils of coils of coils! This allows the 3 billion base pairs in each cell to fit into a space just 6 microns across.
If you stretched the DNA in one cell all the way out, it would be about 2m long and all the DNA in all your cells put together would be about twice the diameter of the Solar System.” (Please note that I do NOT recommend trying this at home without adult supervision! ;>)
The task of simply finding “gene #11,347” when more of “protein #11,347” is needed is daunting. Come to think of it, just knowing when and how much of “protein #11,347” is needed is an impressive high-level management problem! Each gene has a “header” at the start end. I guess that you could think of it as functioning as a bar code containing some equivalent of “#11,347”. Think of going out with a scanner to find it.
The familiar tidy view of the chromosome is actually only seen when the cell is preparing to divide. During “normal business hours” the chromosomes appear much less organized. There is a reason, of course. Each cell type has a specific set of genes that are relevant and necessary for it to function. Tongue in cheek, I often point out that your liver needs the ability to produce bile. On the other hand, your brain cells have no need to spew out bile — unless you are a Hollywood movie script writer! To expedite normal cell operation, genes that are not needed are “switched off”, while necessary genes are “switched on”.
But first just consider the problem of “DNA Management”. Every cell contains a complete copy of your genome. That DNA has a total length around 6 feet. To grasp the problem more clearly, lets “magnify” your DNA up to the size of old-fashioned telephone handset cords, to represent the DNA double helix. At that size, your DNA would be about 10,000 km long (about 6,000 miles!). Now dump that into a super-sized Olympic swimming pool and jump in to find “gene #11,347”. By the way, be careful not to break or get the DNA tangled.
Figure 2. The “Tangled Phone Cord” analogy.
At the other extreme, “gene #11,347” is hopelessly buried deep within the “coil of coils of coils of coils of coils” in Figure 1.
The cell operates at an optimum intermediate level, also documented in Figure 1. But by now it should not come as much of a surprise that “switching” a gene on or off is much more complex than just a switch. One such switch is called “methylation”. A methyl group (CH3) is added to cytosine DNA nucleotide to suppress copying (“switch off”), while “demethylation” tend to make it more readable. The chromatin loop containing the gene has to be “fished out” from the chromosome so that it can be read. There are “Scotch sticky flags” that are attached to the histone “spools” to draw attention to the location of the “bar code strip” for fast reading.
The ultimate challenge comes at the time of mitosis (cell division). Everything in the cell must be duplicated in order to make two complete daughter cells.
Second, everything must be sorted so that both daughter cells actually end up with a complete complement of “parts”. This is critically true of the chromosomal genetic information: both daughter cells must end up with a complete complement of chromosomes:
The ribosome is much like a “computer-controlled machine”. It “reads” the mRNA strand in three-base “words” called codons. Each codon specifies one of the 20 amino acids used by living organisms to build proteins. The ribosome then assembles a growing string of amino acids until it arrives at a “stop” instruction (a stop codon). It then releases the finished amino acid chain and the mRNA strand. Another ribosome soon picks up the mRNA strand and makes another protein string until the mRNA string “wears out” or is broken down.
This description is, of course, vastly over-simplified. You should learn to expect that nothing in the cell is “as simple as it seems” — even when it already seems utterly baffling!
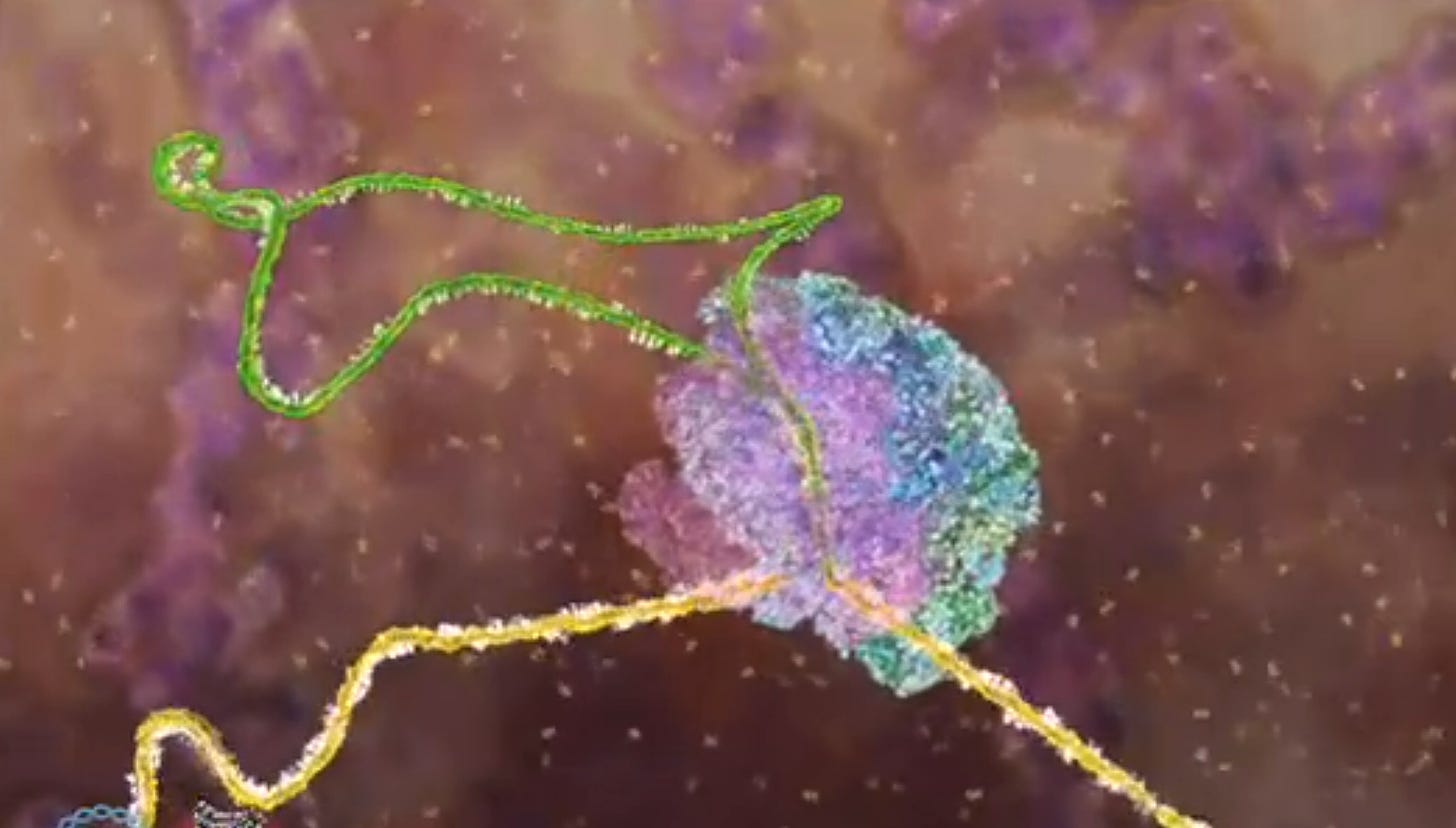
For example, it was initially thought that the coding in a gene had a straightforward 1:1 relationship to the finished product. It was soon discovered that the genes seemed rather fragmented. There were “non-functional” sections called "introns" interspersed throughout the gene that did not end up as part of the finished protein. After the gene is transcribed to a strand of messenger RNA (mRNA), these superfluous sections must be snipped out, and the gaps spliced together[1]. This is performed by spliceosomes. To view an amazing animated video of the process in real time, go to:
https://www.dnalc.org/view/16933-3D-Animation-of-DNA-to-RNA-to-Protein.html .
Here is a simplified graphic of the overall processing of a gene:
Fig. 4 Introns and exons.
This in itself is a startling discovery. This seems to add a lot of "unnecessary" complexity (with potential for serious errors) to the transcription process. But explaining the origin of this gene splicing is particularly problematic. First, there has to be an overall plan. Second, there must be an organized procedure for implementation. Third, this required a group of specialized protein machines which can carry out some complex and critical operations[1]. Fourth, these all had to come together and work flawlessly. The entire evolutionary process could have been completely derailed at any stage of gene splicing.
It was initially unclear whether these introns were simply useless "filler material", but they actually contain additional instructions and functions soon to be discovered which guide and direct the splicing operations. And here is an interesting twist: it seems that the spliceosomes must somehow “read” the exons as they are clipping them out, and somehow “remember” the special instructions before discarding them. Otherwise, they would be “discarding the instructions”. I am unsure how that works, but “I am here to talk about it”, so apparently it does…
But it gets even more complex. Researchers discovered that there are actually overlapping genes. This is accomplished as shown below. Here, three hypothetical proteins share the same "framework" structure that supports the active site of each protein. By selecting which "optional" exons will be copied, several different "final products" can be stored in the same gene. The less-critical “supporting structure” design is re-used for making several different proteins. In other cases, this supporting structure has to be "bent" slightly to put pressure on the active site to fine-tune it for absolute maximum performance. Such a protein cannot use a generic "stock" framework unless alternate "modification exons" are specified. But again, in all cases the spliceosomes must remember the exon instructions that they clipped out and discarded.
The spliceosomes also add a “header” to properly orient the mRNA and a “Poly-A” tail. The tail might be 20 to several hundred bases long. A longer Poly-A tail takes longer to erode and degrade, thus that mRNA produces more protein copies.
Only after selective splicing is the mRNA ready for production. The mRNA is then ushered out of the nucleus, where ribosomes pick up the mRNA strands and start assembling the specified chains of amino acids. These immediately start to self-fold into a variety of basic sub-structures, such as the alpha helix, the beta pleated sheet, etc.
The alpha helix and other structures will fold into their final three-dimensional shape to complete the process in the next section.
[1] p 103: Much of this discussion comes from The Cell's Design by Fazale Rana (Colorado Rapids, Baker Books, 2008). Available from Amazon. Page number references are from this book.