Decoding the Genetic Code - Part 3
The Protein Assembly Shop Edited August 19, 2022
In Part 1, I discussed the basics of the Universal Genetic Code. It is amazingly optimized, resistant to errors, and highly flexible.
In Part 2, we examined the complex process of finding the gene which is needed, making an accurate copy, meticulously preprocessing the mRNA, and passing it on to “manufacturing”.
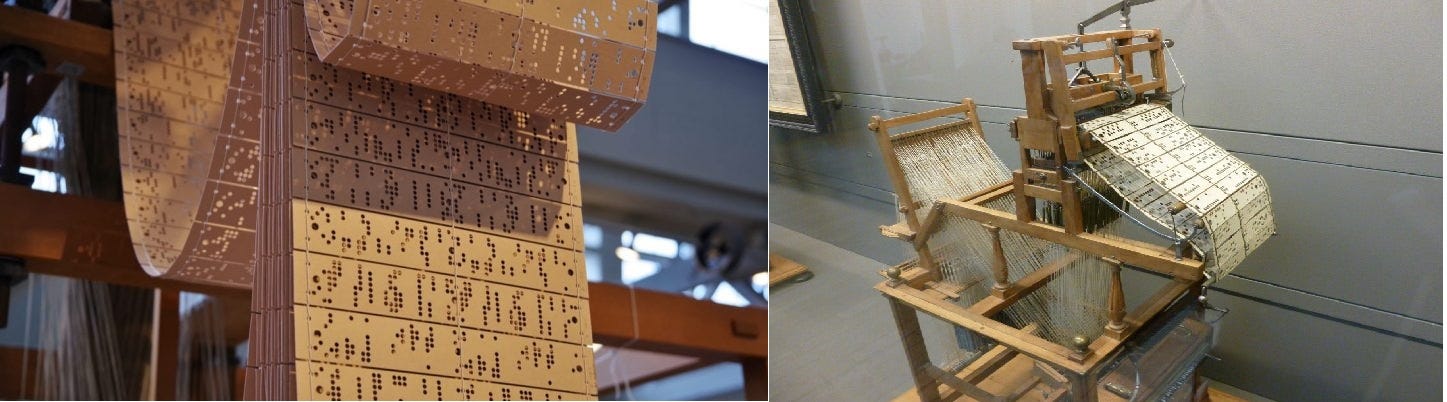
When Joseph-Marie Jacquard, a French weaver and merchant, patented his invention in 1804, he revolutionized how patterned cloth could be woven. His Jacquard machine made it possible for complex and detailed patterns to be manufactured automatically.
The punched cards contain the pattern to be woven. Each hole specifies whether the next thread being woven passes over or under, thus generating the pattern. After weaving run of a selected pattern, the punched cards were returned to a rack for future use.
Some have flagged this as “the start of the computer age” since binary data (the holes in the cards” was directly used to control a machine. This idea later evolved into the IBM punched cards and the punched paper tapes of the early computer age.
But it also serves as a good illustration of the way that the cell produces building materials (proteins) and reproduces.
We discussed in Part 2 how the “punched paper tapes” (mRNA) are copied from the DNA and transferred to the “shop floor”, the cell cytoplasm. The “machine” that reads them is the ribosome, and it is indeed an amazing production machine.
The ribosome is actually made up of many separate parts, like a typical machine. There are about fifty individual RNA and DNA protein “parts” that all self-assemble into a functional “duplicating machine”. Every one of those parts is precisely shaped so that they only fit together one way, and when they do come together, they “snap” permanently into place. So not only are the shapes precisely matched, but the surface electrical charges form a kind of “velcro” that holds the whole machine together. But the parts also have to be free to move, just like an industrial machine, and those movements need tiny “motors” to move them energetically. This energy is supplied by ATP, which powers most other activity in the cells. There are roughly one million atoms in a ribosome, at the finest detail. It is a very complex machine.
The ribosome takes in a floating mRNA string and reads past the header to the START codon, where it begins assembling the amino acid string. For each “3-letter” codon, it selects the specified amino acid and attaches it to the growing string. When it comes to a STOP codon, there is no specified amino acid, so that terminates the string. As the mRNA string floats away, it is picked up by another ribosome which manufactures another protein molecule. This continues until the mRNA wears out and breaks down. So instead of “returning the template to the storage rack”, the cell will copy a new mRNA from the library in your DNA as needed.
The Jacquard cards were slats of wood, so they would eventually splinter. A technician would have to “repair the DNA” by replacing that card. The cell has similar functions to repair your DNA.
Fixing Source Code Errors
During the replication of your DNA prior to cell division, errors sometimes occur. Considering the huge number of base-pairs (over 3 billion), a substantial number of errors are inevitable. To understand better why errors are "inevitable", just watch the process of DNA replication in real time:
https://media.hhmi.org/biointeractive/media/DNAi_replicacion_vo2-esp.mp4?download=true (SPANISH)
There is a whole suite of proteins that come to the aid of DNA Synthetase to perform a bewilderingly complex task at what seems like a break-neck speed. Note that this video is in "real time". They sometimes refer to DNA as being "self-replicating". That absolutely is NOT TRUE. It takes a small army of highly complex proteins to perform extremely complex operations to manufacture the duplicated DNA.
But if DNA can’t self-replicate, that presents a serious problem. Very early in the evolutionary development — before the small army of highly complex proteins to perform extremely complex operations even existed — DNA needed to replicate for the first time. The genes for DNA Synthetase, gyrase, ribosomes, (and a host of others) was still “under development” inside the unreplicated DNA. It needed ribosomes to manufacture all of those proteins, but the “first” ribosome needed a previously existing ribosome to manufacture the “first” one. This is vastly more complex than a”chicken and the egg” problem.
This leads to another marvel that is still not well understood. There is error correction at work on the "source code": your DNA itself. A mutation in the DNA not only affects the next protein to be manufactured, but all subsequent copies as well. One research group has discovered over 700 proteins whose function is to traverse the length of your genome looking for errors. When it finds an error, it snips out that segment and corrects the error. 700 proteins is a major significant fraction (3.5%) of your 20,000 proteins. Now obviously these mere 700 proteins can't "remember" my whole genome to compare and find specific errors. That would at least double the size of the genome if they carried "proof copies"! But since the master copy itself has to be copied, it could contain errors. It is suspected that the flexibility in "spelling" of protein sequences allows for the definition of "acceptable" and "unacceptable" longer sequences. If the gene coding selectively avoids forbidden sequences and leans strongly towards preferred sequences then the "Mr. Fix-It" enzymes (yes, that is what the web site actually named them ;>) do not need to know the specific "original code", but needs only to correct "unacceptable pattern" errors. This is much like parity coding or the error metric used in FEC (Forward Error Correction coding) in high speed digital transmissions. Thus there is evidently a high level of quality control at work at every level of the cell.
With this text that I am typing the “spell checker” can verify most of the common words that I might misspell. However, since this is a technical paper, there are a number of words that the “spell checker” has not yet “learned”. Glancing over the text, “Synthetase ” and “FEC” pop up. The situation is quite different with genetic code. It is also different with numerical data like the 2020 Census. There is no way to “spell check” such data. This is where an analogy between Forward Error Correction coding and the redundancy of the Genetic Code might be useful, because it is the only way that “genetic error correction” seems even remotely possible. I pointed out in Part 1 that “A protein with a length of 100 residues can be "spelled" roughly 3^100 ways. That is 5 x 10^47 ways to "spell" the exact same protein. But even any segment as short as 12 base pairs (4 amino acids) can be "spelled" about 81 ways and any segment of 6 base pairs (just two amino acids) can be written 9 different ways.”
Obviously a simple spelling rule like “i before e, except after c” could not apply to raw genetic code. On the other hand, given the tremendous “flexibility” in spelling proteins, it is not hard to imagine that, if the spelling could be “adjusted”, it might be entirely possible to accommodate such a rule. Given that there are 700 such “spell checker” proteins, there could actually be 700 “simple spelling rules” which collectively are fully capable of collecting errors without having an actual “master original copy”.
Admittedly, this is armchair speculation. Better yet, lets call this “scientific theory”. (Sometimes the two are indistinguishable.) This is an area where some creative thinking is sorely needed.
Those “Mr. Fix-it proteins” (700 of them!) are a case in point. The Darwinist explanation is that they “evolved the ability to correct errors”. Have they thought through the implications? Consider the very first “evolving error-correction protein machine”. It is equipped with some means for “crawling” the length of your DNA and reading your genetic code. It is somehow able to recognize “improper” or “forbidden” code sequences. It is equipped with surgical instruments to “snip out” the bad code and then “glue in” the correct code. The problem of course is that it is evolving. You could call it “on-the-job-training”. It will take a while for it to figure out what is “erroneous code” and what isn’t”. This is like having a first-grader doing the spell-checking on your document! Meanwhile, this “evolving” error-fixer will be creating chaos. And remember next that there are 699 apprentices still to come. The best description that I can think of for this Darwinian Evolution is quite frankly “DNA Cole Slaw”. Before they “learn” how to do it right, random, unguided trial-and-error would be catastrophic. This is just one of those functions which cannot possibly evolve.
All of this discussion of course assumes that that the “protein re-spelling scheme” was already in place. Perhaps they could theorize that the rules “evolved” one at a time.
These questions pose a fundamental threat to Darwinian thinking. Error correction implies a Higher Intelligence. This is fairly obvious. But beyond that, it tends to be detrimental to evolution. Many random mutations will also violate one or more of the “700 spelling rules” and will be corrected and “un-evolved”. The Darwinist would prefer to see free and unfettered “experimentation”. But “evolving” proteins need to meet the further constraint of correct spelling, otherwise they can’t be protected, or they may be eliminated.
Gaining a fundamental understanding of how DNA error correction works is a fundamentally critical endeavor. Error correction is found in even the earliest life forms and without error correction they would not have survived. How did highly sophisticated functions develop in the “simplest, oldest life forms”? There are many genetic disorders that involve failures of certain error corrections. Unfortunately, as I have pointed out, the evolution of many of these functions are “theoretically impossible”, and thus the Darwinist just doesn’t venture a guess at understanding them. For the Creationist who believes in Intelligent Design, it is not at all difficult to think seriously about these questions and suggest possible areas for investigation. I might say that we have an “unfair advantage” in understanding these matters.
It is one thing to discuss and describe these high-level functions. But implementing them in tiny "dumb" proteins is a whole other story. These are tiny protein complexes which bind to the DNA double helix. They then "read" (as by “Braille”, perhaps? ;>) the DNA code. They are able to "recognize" improper patterns and react to them by performing delicate and complex surgery. In principle, we could build highly complex computer-controlled robots which could be programmed to perform equivalent functions, (on a larger scale, or course!) but even that would require an immense amount of intelligence.
Inventing these "purely mechanical" proteins would be vastly more difficult. It is amusing to consider an "equivalent machine". It must wrap around the DNA double helix to access the code through the spiral slot. It would have a motor that propels it along. It has "fingers" that "feel" the code. When they feel a match, they drop down like the tumblers in a lock. This releases a latch and starts some "gears and cams" rotating. This particular "MR. Fix-It" has a specific pre-programmed task. It opens the DNA helix, does what is needed to correct the error in this case, then returns the DNA to normal. All of these actions are energy deficient and probably require an ATP "battery" to dump its charge to ratchet a motor protein. But this machine has to recognize the particular type of error and take appropriate action. (Perhaps that is why there are 700.)
For simple base-pair mismatches (they should always be A-T or C-G) it must know which strand is the master copy and which is the "back-up copy" (the complement). Correcting the master copy to match the complement would make the error permanent, since this is presumably an error that occurred during DNA replication.
Other cases escalate in complexity. So an individual Mr. Fix-it may find a suspected error, but it moves on because that problem is not within its "skill set". When the proper Mr. Fix-it recognizes "my kind of problem" it then goes into action, which may involve a number of complex and delicate steps. (see graphics below) You must realize that a malfunctioning Mr. Fix-it is a major fiasco. "On the job training" of an "evolving" Mr. Fix-it would generally result in "DNA cole slaw".
Again, most of these actions are energy deficient and probably require ATP "batteries" to dump their charge to ratchet a motor protein. Or perhaps you might think of these motors as "internal combustion engines"... When ATP releases the third phosphate, it does so energetically. But you must somehow know how to harness that energy. The motor proteins have a "slot" for ATP which will strategically direct that energy precisely where it is needed to do specific work efficiently. They also have a "spark plug" or "firing pin" that controls the energy release. And of course they must "exhaust" the spent ADP and "load-and-lock" a new ATP. Since the great majority of functions use ATP for energy, the entire metabolic pathway must be implemented simultaneously. ATP is not useful unless complex machines exist that are specifically designed to use that energy, but complex motors and processes that use it will not develop unless ATP already exists. These "chicken or the egg" scenarios are extremely difficult to explain is evolutionary terms, and yet they are more the rule than the exception in biology.
There is yet another very troubling aspect of this for the evolutionists. In Richard Lenski's experiments[1] at the University of Michigan, he has been growing E. Coli cultures for a quarter century. that is over 60,000 generations being acted on relentlessly by Darwinian selection. In several cultures, this error-correction mechanism suffered serious damage. Suddenly the mutation rate of that culture jumped up a hundred-fold. That seems indicate that these amazing error-correction enzymes are 99% effective in correcting mutation errors -- including those potentially "helpful" mutations that evolutionists are looking for! If so, then the "Mr. Fixit" proteins are slowing down Darwinism by a factor of 100. This was NOT what they are looking for!
[1] See the chapter in Darwin Devolves by Michael Behe.
The escalating levels of damage needing repair, combined with the increasing complexity of repair operations lend rapidly increasing weight to the argument that this all requires intelligent intervention.
Since there are 700 "Mr. Fix-it" proteins, it is obviously beyond the scope of this paper (or anyone's paper) to detail all of these actions. Just be assured that the Quality Control is in place, and it is breath-taking!
The "Tiny Big Picture"
We have been examining details on the molecular level. True, proteins are very large molecules, but they are still tiny. There are many millions of protein molecules in a single cell. If we zoom out to see one whole cell, a huge higher order of complexity comes into view.
The cell has an outer "skin" (the cell wall) which keeps the cell contents and chemistry separate from its environment. There is an inner membrane that encloses the nucleus where the DNA resides.
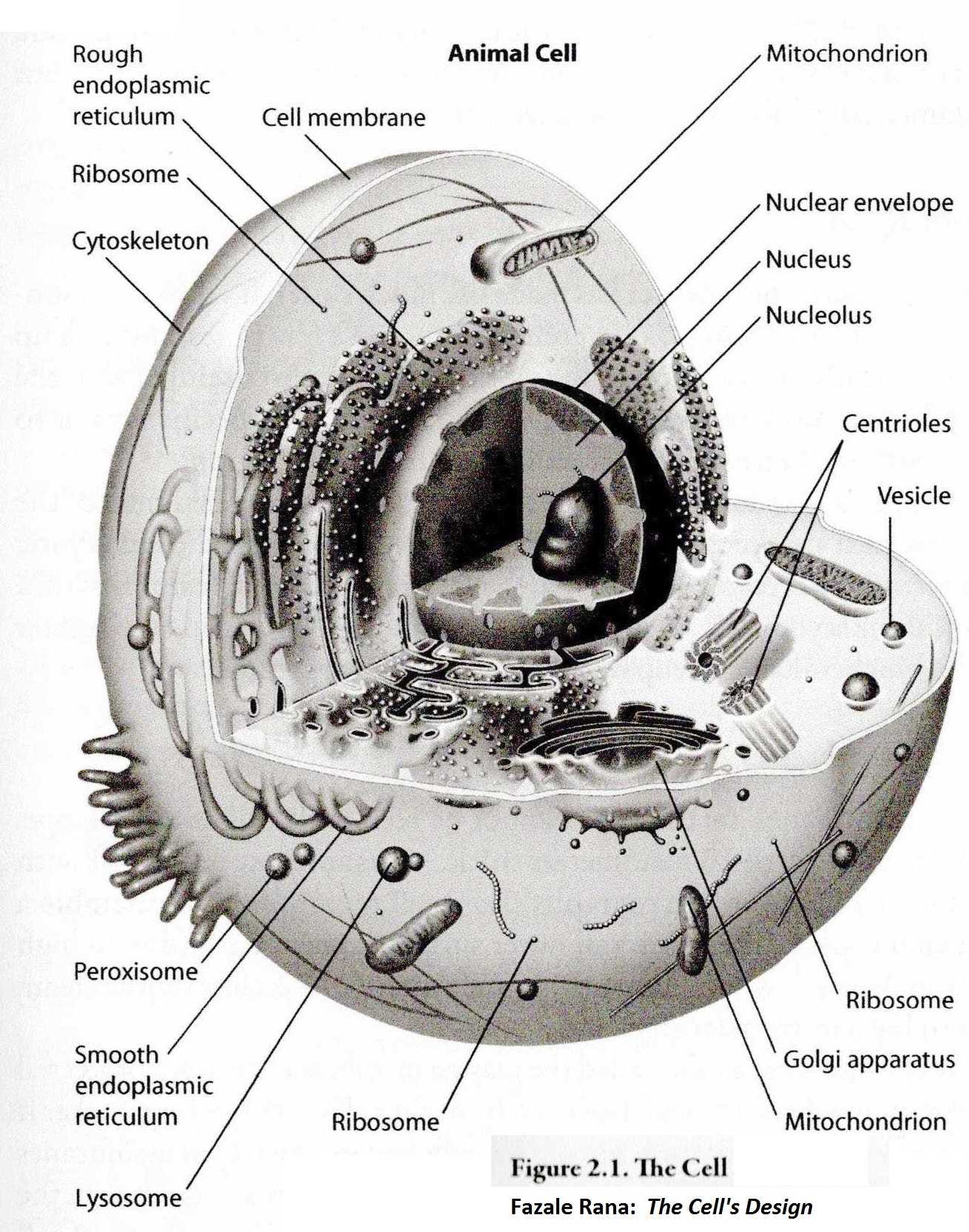
The most prominent substructure within the nucleus is the nucleolus (see figure below), which is the site of rRNA transcription and processing, and of ribosome assembly. Cells require large numbers of ribosomes to meet their needs for protein synthesis. Every protein in the cell was manufactured by a ribosome. Actively growing mammalian cells, for example, contain 5 million to 10 million ribosomes. A whole new set must be synthesized each time the cell divides. These in turn manufacture the hundreds of millions of proteins in a single cell, one protein molecule at a time. The ribosome is literally a tiny CNC machine that "reads" the mRNA copy of a gene and assembles a protein, one amino acid at a time. The nucleolus is a ribosome production factory, designed to fulfill the need for large-scale production of rRNAs and assembly of the ribosomal sub-units. Think of it: this is a dedicated production facility with some 5,000 ribosome CNCs that is constantly manufacturing new ribosome CNCs to keep the 5 - 10 million ribosome CNCs running that build the rest of the cell. One ribosome is made up of about 80 protein "parts" that fit together to make up the complete machine. There are about one million atoms in one ribosome. In the drawing below there are several ribosomes labeled. There are actually more than 5,000,000 in many cells.
Consider the implications. Somehow, the individual cell must control and regulate exactly how many of every protein to make. None can be ignored and non can be over-produced. When the cell prepares to divide (mitosis) it must call up a complete "bill of materials" to start building all of the parts needed for a complete second cell. A certain minimum must be met before cell division begins, but perhaps some "details" can be deferred until after division, but even that takes planning. There are a host of special materials (proteins) that are only needed during the division process.
Speaking of that "Bill of Materials", these are contained in the genes. Each gene has the "instructions" for making one protein. But how does the cell keep track?
Your DNA in one cell is about 2 nanometers in diameter, and 2 meters (six feet) long! That is one billion times longer than the diameter. If your DNA were the diameter of a coiled telephone cord, it would be 6,200 miles (10km) long. How does the cell "index" all of the 20,000 genes and how does it quickly find the exact gene that needs to go to production? How does it know when it has made "enough", or how does it know it is becoming deficient in yet another? And how does it "plan ahead" for cell division?
For years we have heard about DNA being transcribed into messenger RNA (mRNA) which is then used to control the production of proteins. But this is only a couple of percent of the genome. I recall during the Human Genome Project (1990-2003) there was a great deal of emphasis on "genes". These, they explained often, were the instructions for actually building proteins -- hence, your body. But these constituted only about 20% of the genome. This was later revised downward to "less than 10%", and eventually to "less than 5%". At that point a reporter asked the obvious question: "Then what is the other 95%?"
Well, the rest was just "Junk DNA". It doesn't actually make anything. It was just the "leftover scratch-pad of evolution". It probably contained obsolete evolutionary experiments and "earlier models".
In the November 2003 Scientific American[1], Michel Georges, a geneticist at the University of Liege in Belgium is quoted as saying that many of these sequences are clearly functional, even though they do not code for any protein. John S. Mattick, director of the Institute for Molecular Bioscience at the University of Queensland in Brisbane, Australia, admits, “I think this will come to be a classic story of orthodoxy (evolution – dwh) derailing objective analysis of the facts, in this case for a quarter of a century. The failure to recognize the full implications of this – particularly the possibility that the intervening non-coding sequences may be transmitting parallel information in the form of RNA molecules – may well go down as one of the biggest mistakes in the history of molecular biology.” (I must point out that it wasn’t Creationism that set back progress 25 years, it was Evolution! The "Junk DNA" concept fit so perfectly with evolutionary theory that this explanation was almost universally accepted without further thought.)
I would think that it should have been obvious to any thinking person that the genes only make proteins. The genes are thus just the equivalent of the "Bill of Materials" for my body (with the recipe for those materials). Where was the Assembly Manual? Where was the Repair Manual? Where was the Operating Manual? ...just to scratch the surface! It should have been obvious that these had to be in the "Junk DNA".
It has now been documented that over 90% of the genome gets copied into ncRNA. That is short for "non-coding RNA", formerly known as "Junk DNA"[2].
I would suggest that you go to watch a few of the stunning movies of microscopic embryo development[3]. Somewhere in the "Junk DNA" there is the complete choreography for the growth and assembly of the embryo. It also contains the enormously complex catalog and indexing system for keeping track of all of the 20,000 genes, and the "search engines" that go out and find them.
So, as difficult as that was, "finding the genes" didn't even scratch the surface of the complexity in the genome. But evolutionary theory consistently and grossly underestimates the complexity. The truth is that it has to keep that estimate as low as possible, because evolution can't explain 1% of the complexity we have discovered even now. The explanation for just about everything seems to be, "Isn't it wonderful what a million years of evolution can accomplish?" No, I don't frankly believe it can...
Belief in a Creator confers a tremendous advantage on a scientist. He can (and should) anticipate complexity that is literally unexplainable. Also, instead of wasting time trying to conjure up a story of "how this evolved from that", he can get directly to the real questions: "Why was that designed this way?" and "What is the purpose?"
Frankly, I don't think that even, "Isn't it wonderful what a billion years of evolution can accomplish?" will work any more[4].
[1] “The Unseen Genome: Gems among the Junk”, by W. Wayt Gibbs, Scientific American, November 2003, pp.46-53.
[2] https://www.news-medical.net/news/20180411/Researchers-discover-how-junk-DNA-plays-key-role-in-holding-the-genome-together.aspx
http://www.digital-embryo.org/
[4] Decoding the Genetic Code210726.doc